Metatranscriptomic and Thermodynamic Insights into Medium-Chain Fatty Acid Production Using an Anaerobic Microbiome
M.J. Scarborough et al. (2018). Metatranscriptomic and Thermodynamic Insights into Medium-Chain Fatty Acid Production Using an Anaerobic Microbiome. mSystems 3:e00221-18
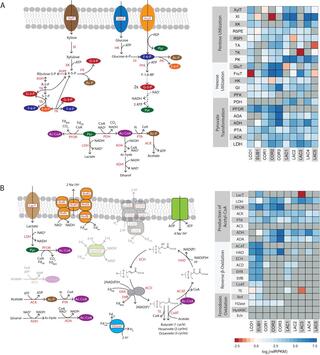
Biomanufacturing from renewable feedstocks can offset fossil fuel-based chemical production. One potential biomanufacturing strategy is production of medium-chain fatty acids (MCFA) from organic feedstocks using either pure cultures or microbiomes. While the set of microbes in a microbiome can often metabolize organic materials of greater diversity than a single species can and while the role of specific species may be known, knowledge of the carbon and energy flow within and between organisms in MCFA-producing microbiomes is only now starting to emerge. Here, we integrated metagenomic, metatranscriptomic, and thermodynamic analyses to predict and characterize the metabolic network of an anaerobic microbiome producing MCFA from organic matter derived from lignocellulosic ethanol fermentation conversion residue. A total of 37 high-quality (>80% complete, <10% contamination) metagenome-assembled genomes (MAGs) were recovered from the microbiome, and metabolic reconstruction of the 10 most abundant MAGs was performed. Metabolic reconstruction combined with metatranscriptomic analysis predicted that organisms affiliated with Lactobacillus and Coriobacteriaceae would degrade carbohydrates and ferment sugars to lactate and acetate. Lachnospiraceae- and Eubacteriaceae-affiliated organisms were predicted to transform these fermentation products to MCFA. Thermodynamic analyses identified conditions under which H2 is expected to be either produced or consumed, suggesting a potential role of H2 partial pressure in MCFA production. From an integrated systems analysis perspective, we propose that MCFA production could be improved if microbiomes were engineered to use homofermentative instead of heterofermentative Lactobacillus and if MCFA-producing organisms were engineered to preferentially use a thioesterase instead of a coenzyme A (CoA) transferase as the terminal enzyme in reverse β-oxidation.
Raw DNA and cDNA read data can be found on the National Center for Biotechnology Information (NCBI) website under BioProject accession no. PRJNA418244. Sequencing reads are available through the following NCBI sequencing read archive (SRA) accession numbers: DNA day 12, SRR6292603; DNA day 48, SRR6292602; DNA day 84, SRR6292605; DNA day 96, SRR6292604; DNA day 120, SRR6292607; RNA day 96 A, SRR6292606; RNA day 96 B, SRR6292609; RNA day 96 C, SRR6292608.