Research Highlights

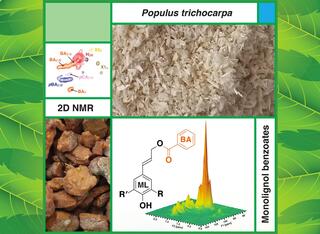
Great Lakes Bioenergy researchers and collaborators engineered softwoods to incorporate a key feature of hardwoods. The resulting pine (shown here) processes more easily into pulp and paper.
Great Lakes Bioenergy research consistently results in new discoveries and new technologies. Here, we highlight high-impact research from all three of our research areas.