Research Highlights

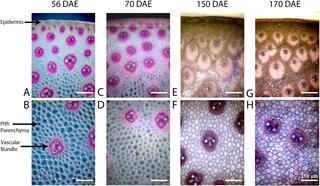
Great Lakes Bioenergy researchers and collaborators engineered softwoods to incorporate a key feature of hardwoods. The resulting pine (shown here) processes more easily into pulp and paper.
Great Lakes Bioenergy research consistently results in new discoveries and new technologies. Here, we highlight high-impact research from all three of our research areas.
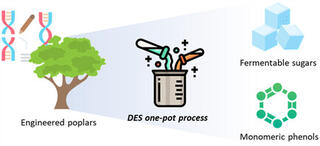
Engineered poplar enhances efficiency of one-pot biorefinery process
Lignocellulosic biomass is an abundant renewable resource that could provide fuels and chemicals for a bioeconomy, but the complexity of plant cell wall structure limits its use. Treatment with environmentally safe and inexpensive deep eutectic solvents (DES) is a promising technique for recovering the three main cell wall polymers: cellulose, hemicellulose, and lignin. Research has also focused on genetic engineering to produce lignin that is easier to deconstruct. Here, scientists evaluated three engineered poplar strains with distinct lignin modifications to assess how this tailored lignin chemistry influences deconstruction and sugar release in DES-based conversion.
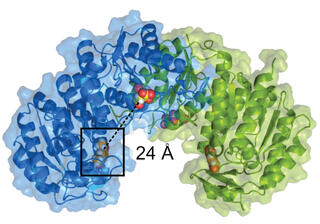
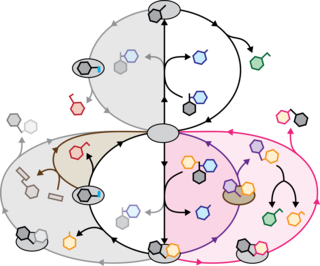
Novosphingobium aromaticivorans enzyme enables funneling of additional aromatics
The plant polymer lignin is an abundant renewable source of aromatics. Chemical depolymerization yields mixtures of aromatics, including acetovanillone, a vanillin derivative with an acetyl side chain. Not all microbes that can generate chemicals from aromatics can metabolize acetovanillone, which can represent up to 10% of the aromatic monomers in deconstructed biomass. Here scientists identified a single amino acid change in a previously uncharacterized protein that is necessary and sufficient for N. aromaticivorans growth with acetovanillone as the sole organic carbon source.
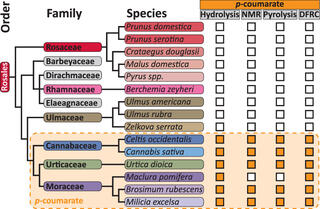
p-Coumaroylated Lignins Found in Three Rosales Families
The complex carbon-rich plant structure lignin can be broken down to make aviation fuel, plastics, and other commercial products. During chemical deconstruction of lignocellulosic biomass, the presence of phenolic pendant groups such as p-coumarate can improve the efficiency of deconstruction or lead to toxins that inhibit microbial fermentation of plant sugars. Though previously thought only to be in commelinid monocots, the list of pCA-containing eudicots is growing.
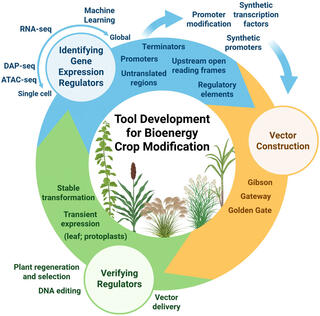
Mapping advances and bottlenecks on the path to engineering bioenergy crops
This work reviews the current landscape of plant genetics through the lens of bioenergy crops to present a roadmap for using DNA-based tools to engineer improvements.
Analysis reveals causes of threefold increase in bioenergy sorghum stem density
Bioenergy sorghum is a drought-tolerant grass adapted low-productivity lands that promotes soil carbon stocks. An extended vegetative growth phase and long growing season produce 4-5 meter stems that account for about 80% of shoot biomass. During a typical growing season stem density increases significantly following stem internode growth. This study offers new insights on what causes it.
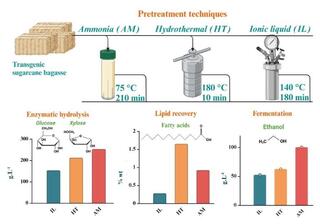
Evaluating the Industrial Potential of Emerging Biomass Pretreatment Technologies in Bioethanol Production
This research enhances the selection and validation of upstream processing methods for lignocellulosic biomass pipeline processing, which improves economic and environmental outcomes for biorefineries, and shows the first comparison of pretreatment technique viability for oilcane.
Analysis of poplar hydrogenolysis reveals new pathways and products
Grasses and some tree species have naturally γ-acylated lignins. Poplar lignins have para-hydroxybenzoate groups on 1-15% of syringyl subunits. During hydrogenolysis, it is generally assumed that p-hydroxybenzoate is cleaved before the deacylated lignin is depolymerized. Here, scientists showed how the presence of a γ-acylated group alters the product portfolio produced by hydrogenolysis with palladium on carbon (Pd/C) as the catalyst.
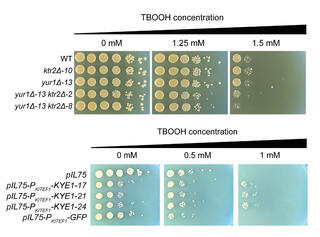
Machine learning reveals genes impacting oxidative stress resistance across yeasts
Researchers characterized variation in ROS resistance across the ancient subphylum Saccharomycotina and used machine learning to identify gene families whose sizes were predictive of ROS resistance. The most predictive features were enriched in gene families related to cell wall organization and included two reductase gene families.
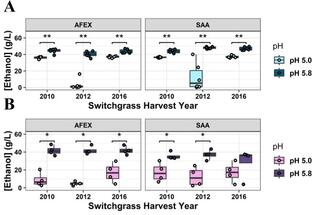
pH adjustment increases biofuel production from drought switchgrass hydrolysate
Switchgrass grown during drought conditions has high levels of osmoprotectant sugars and saponins that inhibit microbial conversion to biofuels. This experiment explored whether the inhibitory effect was specific to ammonia fiber expansion (AFEX) pretreatment and whether it could be alleviated by raising the pH of the hydrolysate.
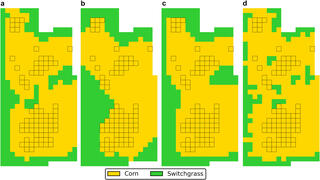
Model balances profit, biodiversity, and ecosystem services to guide bioenergy crop layout
Great Lakes Bioenergy Research Center scientists developed a mixed-integer quadratically constrained program to optimize the layout of a field-scale cropland considering economic, biodiversity, greenhouse gas emissions, and water quality objectives. Decision variables include spatially varying fertilization in addition to crop establishment location. The model also accounts for biodiversity effects of core area and edges between crops.