Research Highlights

Great Lakes Bioenergy researchers and collaborators engineered softwoods to incorporate a key feature of hardwoods. The resulting pine (shown here) processes more easily into pulp and paper.
Great Lakes Bioenergy research consistently results in new discoveries and new technologies. Here, we highlight high-impact research from all three of our research areas.
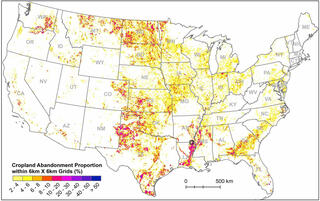
Atlas pinpoints abandoned croplands for potential biofuel production
Researchers used existing land cover data sets to train a computer algorithm to classify land use patterns in satellite imagery. These classifiers were then used to map cropland annually from 1986 to 2018 at a 30-meter resolution. Researchers then identified the location and time of abandonment using a moving temporal window for each pixel and estimated the accuracy against visually interpreted sample locations and publicly available datasets.
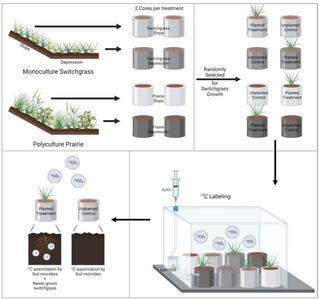
Prairie soil clings to carbon on slopes
Belowground carbon content was higher in the switchgrass grown in prairie soil. Switchgrass grown in prairie soils situated on slopes also had higher biomass carbon in both the above- and belowground plant growth as compared to switchgrass grown in the prairie soil of depressions. Prairie systems consistently outcompete monoculture systems in plant diversity, carbon content, and microbial carbon content.
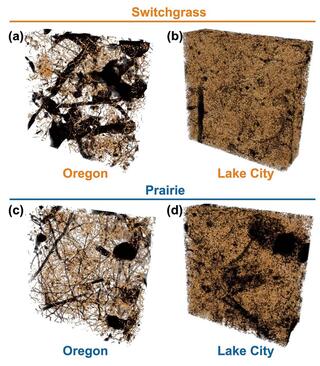
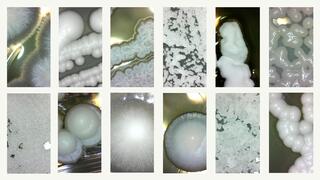
Detritusphere microenvironments play an important role in carbon dynamics
Findings suggest that decomposition dynamics of plant residues is a combined effect of vegetative history, in part through its impact on microbial communities, the chemical and physical characteristics of plant residue, and soil pore structure, which together create temporally dynamic micro-environmental conditions that influence decomposition.
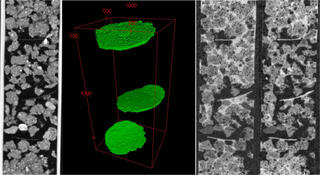
Prairie systems facilitate rapid uptake of carbon
The study assessed soil carbon gains across a variety of marginal soils at experimental sites in Michigan and Wisconsin. Cropping systems were randomly assigned to plots within each of the six unfertilized and untilled sites. Researchers used X-ray computed microtomography to analyze pore structures in harvested soil cores and loose soil surrounding the cores.
Modified poplar lignin eases degradation, production of commodity chemicals
Great Lakes Bioenergy Research Center scientists have patented technology that can be used to make plants with modified lignin amenable to degradation and production of commodity chemicals used in pharmaceutical drugs and cosmetics, potentially enhancing the value to biorefineries.
Efficient production of p-hydroxybenzamide from poplar biomass
The newly patented process uses water as the solvent, no chromatography, inexpensive reagents, no protecting groups, and scalable technology. Modifications of this process have the potential to produce an array of chemical building blocks for the manufacturing materials and chemical products like plastics, surfactants, pigments, and pharmaceuticals.
Genomic factors shape carbon and nitrogen metabolic niche breadth in yeasts
Researchers assembled genomic, metabolic, and ecological data for 1,154 yeast strains, which represent nearly all known species in the subphylum Saccharomycotina, and quantified variation in genome sequence, isolation environment, and carbon and nitrogen metabolism.
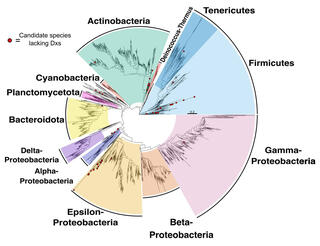
Study reveals potential detours to bottlenecks in microbial terpenoid production
Researchers surveyed a database of 4,400 diverse bacterial genomes, using comparative genomics to identify orthologs of MEP and mevalonate pathway genes. In particular, they looked for alternatives to circumvent the IspG and IspH enzymes, which pose known engineering constraints, or to Dxs, for which some alternatives exist.
Raffinose oligosaccharides support sorghum productivity and resilience
Researchers identified the sorghum genes responsible for making and breaking down raffinose family oligosaccharides (RFOs) and analyzed their activities in different leaf and stem cell types. The results indicate RFOs are produced in leaf cells responsible for photosynthesis and broken down in veins, releasing sucrose where it can be transported throughout the plant. This suggests that RFOs improve sucrose distribution by enhancing short-distance movement within organs.
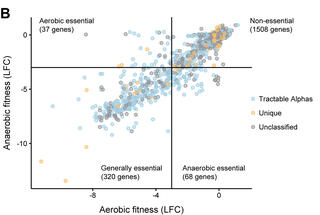
The genetics of aerotolerant growth in Zymomonas mobilis
With fewer than 2,000 protein-encoding genes, Zymomonas mobilis has fewer than half the genes of its closest relatives, is good at converting sugar into ethanol, and able to thrive with or without oxygen. This combination of simplicity, efficiency, and versatility make Z. mobilis a promising model for understanding biology and a potential industrial workhorse. Yet the genes required for growth in various conditions have not been well studied.