Research Highlights

Great Lakes Bioenergy researchers and collaborators engineered softwoods to incorporate a key feature of hardwoods. The resulting pine (shown here) processes more easily into pulp and paper.
Great Lakes Bioenergy research consistently results in new discoveries and new technologies. Here, we highlight high-impact research from all three of our research areas.
Study advances understanding of nodal root bud development in energy sorghum
Researchers used microscopy, transcriptome analysis, and gene regulatory network analysis to study nodal root bud initiation and development in bioenergy sorghum hybrids grown at the Texas A&M University Farm. A system of about 175 nodal roots that extend more than 2 meters enable sorghum to absorb water and nutrients while storing carbon in the soil, contributing to the plant's resilience and sustainability as a source of bioenergy.
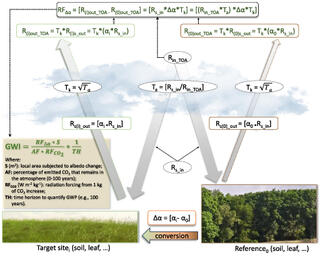
Overlooked Cooling Effects of Albedo
The trapping of heat energy is a significant driver of global climate change. The absorbent nature of materials, such as asphalt, used for infrastructure can lead to an increase in temperature. When determining the value of various crop systems on our shared future, albedo is an often overlooked aspect of the calculation that may seriously impact climate patterns on this planet. The paper shows that ecosystem types, as well as time of day and season, heavily impact the reflectivity of ground cover types, suggesting that new techniques need to be developed if albedo is to be properly considered in carbon credit programs and future policies.
Researchers develop genetic toolkit for engineering Alphaproteobacteria
Researchers developed facile and reliable genetic tools to target essential or other specific genes in the Alphaproteobacteria Novosphingobium aromaticivorans and Rhodobacter sphaeroides, which show promise as organisms that can be engineered to convert plant lignin and sugars into biofuels and commodity chemicals.
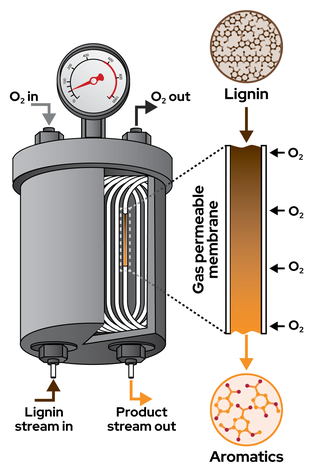
Oxygen-permeable membrane reactor offers scalable approach to lignin depolymerization
Lignin is the world’s largest source of renewable aromatic compounds, but its heterogeneous composition and sequence that varies by source make it difficult to use as a chemical feedstock. Oxidative depolymerization methods generate valuable aromatic products, but such processes are difficult to control because reaction conditions that support depolymerization also contribution to product decomposition.
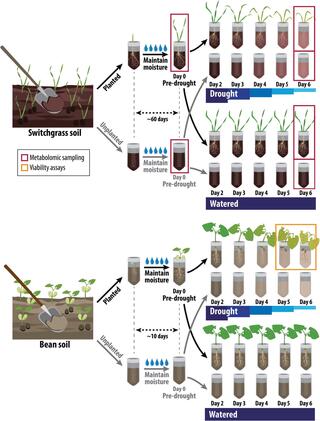
Active root microbe responses to short-term drought are crop specific
The microbiome associated with plant roots, the rhizobiome, can harbor beneficial microbes that alleviate stress, but factors influencing their recruitment are unclear. This greenhouse experiment investigated the impact of short-term drought severity on the recruitment of active bacteria.
Study evaluates potential of genomic prediction to improve switchgrass productivity
Bioenergy crops like switchgrass can pull carbon dioxide from the air and store it in soil while also providing a source of sustainable fuel, but gains in productivity are needed in order to be commercially viable. To improve switchgrass breeding strategies with limited resources by analyzing plant performance across diverse environments, evaluating yield surrogate traits, and developing genomic models to predict biomass yield efficiently.
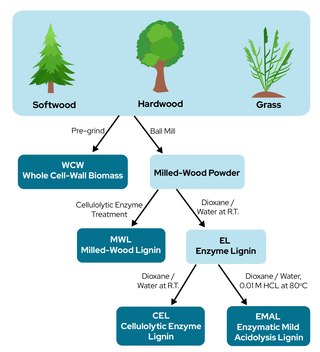
Study provides guidelines for selecting best lignin isolation techniques for research
Lignin, a complex heterogeneous polymer that forms plant biomass structures, is a promising candidate in the biofuel industry because of its aromatic constitution. Although milled wood, enzymatic, cellulolytic enzyme, and enzymatic mild-acidosis lignins have all been studied on behalf of native lignin, they are all the result of different separation strategies and therefore may have differing structures. This study aimed to compare these separation strategies and determine their effects on lignin structure in softwoods, hardwoods, and grasses.
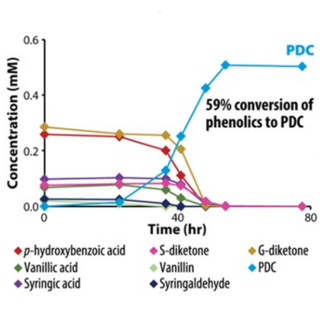
Patent approved for lignin to polyester precursor conversion process
The cost of manufacturing plant-based biofuels has limited competition with cheaper fossil fuels. By engineering one candidate microbe, Novosphingobium aromaticivorans, to funnel heterogeneous mixtures of lignin-derived aromatic compounds to 2-pyrone-4,6-dicarboxylic acid (PDC), a potential bioplastic precursor, this study aimed to increase its potential market value.
Assessing impacts of supply chain variability on centralized cellulosic biorefinery
Cellulosic biofuels derived from crop residues and dedicated energy crops grown on marginal lands are an attractive alternative to liquid fossil fuels. This study seeks to show how annual fluctuations in biomass production impact the economic and environmental performance of biofuels.
Terminal enzymes must be co-optimized to relieve MEP bottleneck in Z. mobilis
Many plants and bacteria use the methylerythritol phosphate (MEP) pathway to synthesize precursors for isoprenoids, a diverse class of hydrocarbons that includes natural products, industrial chemicals and biofuels. This research investigates the function of the oxygen-sensitive enzymes IspG and IspH, known bottlenecks in the MEP pathway of the biofuel-producing microbe Zymomonas mobilis.