Identification of a secondary metabolite gene cluster in budding yeasts with important implications for biofuel production
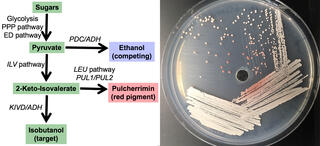
Study of how yeast partition carbon into the metabolite pulcherrimin may provide insight into engineering strategies to boost isobutanol production.
The Science
The iron-binding molecule pulcherrimin was described 65 years ago, but the genes responsible for its production in budding yeasts remained uncharacterized. Genomic comparisons across 90 species of the budding yeast subphylum Saccharomycotina revealed a four-gene cluster associated with pulcherrimin production, and targeted gene disruptions in Kluyveromyces lactis revealed likely functions for each of the genes: two biosynthetic enzymes, a transporter, and a putative transcription factor.
The Impact
Pulcherrimin and isobutanol biosynthesis both partition from the branched chain amino acid (BCAA) biosynthetic pathway, and some yeast strains direct a significant amount of carbon from BCAA biosynthesis into pulcherrimin. Since both pulcherrimin and isobutanol are made from a common pathway this suggests that the metabolic control of high pulcherrimin producers may be harnessed for increased isobutanol production in yeast, wherein most species naturally make limited amounts of isobutanol.
Summary
Pulcherrimin is an iron-binding pigment made by a small number of budding yeast species that is composed of two cyclized and modified leucine molecules. Despite the discovery of pulcherrimin 65 years ago the genes responsible for its biosynthesis remained uncharacterized. Using a comparative genomics approach among 90 genomes from the budding yeast subphylum Saccharomycotina, researchers from the Great Lakes Bioenergy Research Center identified the first yeast secondary metabolite gene cluster and showed that it is responsible for pulcherrimin biosynthesis. Targeted gene disruptions in Kluyveromyces lactis identified putative functions for each of the four genes: two pulcherriminic acid biosynthesis enzymes, a pulcherrimin transporter, and a transcription factor involved in both biosynthesis and transport. The requirement of a functional putative transporter to utilize extracellular pulcherrimin-complexed iron demonstrates that pulcherriminic acid is a siderophore, an iron-chelating compound secreted by microorganisms. This research also characterized and named two genes that previously lacked assigned functions in the fuel-producing model yeast Saccharomyces cerevisiae. The evolution of this gene cluster in budding yeast suggests an ecological role for pulcherrimin akin to other microbial public goods systems. Like isobutanol, pulcherrimin is derived from the BCAA biosynthesis pathway, and some yeasts species are particularly adept at funneling BCAA carbon into pulcherrimin. Thus study of how high-level pulcherrimin producing strains are altered in their metabolic control may inform strategies for increased biofuel production in model organisms.
Contacts (BER PM)
Kent Peters
Program Manager, Office of Biological and Environmental Research
kent.peters@science.doe.gov, 301-903-5549
(PI Contact)
Chris Todd Hittinger
University of Wisconsin–Madison
cthittinger@wisc.edu
Funding
This material is based upon work supported by the U.S. Department of Energy, Office of Science, Office of Biological and Environmental Research under award numbers DE-FC02-07ER64494 and DE-SC0018409, as well as the National Science Foundation under grant numbers DEB-1442148, DEB-1442113, and DEB-1253634. Additional funding was provided by the Pew Charitable Trusts and the Vilas Trust Estate.
Publications
Krause, D. et al, “Functional and evolutionary characterization of a secondary metabolite gene cluster in budding yeasts.” PNAS (2018), DOI: 10.1073/pnas.1806268115.
Related Links
http://www.pnas.org/content/early/2018/10/04/1806268115