GLBRC Data Sets
Highlighted below are a variety of published studies that include data sets that might be of interest to the scientific community and have been deposited in online data repositories. Only data sets published in GLBRC-approved repositories following the FAIR Guiding Principles are highlighted. More information can be found on our guidelines page.
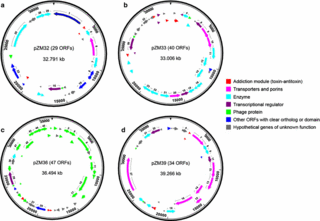
Complete genome sequence and the expression pattern of plasmids of the model ethanologen Zymomonas mobilis ZM4 and its xylose-utilizing derivatives 8b and 2032
S. Yang et al. (2018) Complete genome sequence and the expression pattern of plasmids of the model ethanologen Zymomonas mobilis ZM4 and its xylose-utilizing derivatives 8b and 2032. Biotechnology for Biofuels 11: 125
In this study, we determined the complete chromosome and plasmid sequences of ZM4 and its engineered xylose-utilizing derivatives 2032 and 8b. Compared to previously published and revised ZM4 chromosome sequences, the ZM4 chromosome sequence reported here contains 65 nucleotide sequence variations as well as a 2400-bp insertion.
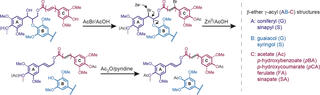
Reductive Cleavage Method for Quantitation of Monolignols and Low‐Abundance Monolignol Conjugates
M. Regner et al. (2018). Reductive Cleavage Method for Quantitation of Monolignols and Low-Abundance Monolignol Conjugates ChemSusChem 11:1600-1605.
As the derivatization followed by reductive cleavage (DFRC) method is particularly effective for structurally characterizing natively acylated lignins, we used an array of synthetic β‐ether γ‐acylated model compounds to determine theoretical yields for all monolignol conjugates currently known to exist in lignin, and we synthesized a new set of deuterated analogs as internal standards for quantification using GC–MS/MS.
The Sorghum bicolor reference genome: improved assembly, gene annotations, a transcriptome atlas, and signatures of genome organization
R.F. Mccormick et al. (2017). The Sorghum bicolor reference genome: improved assembly, gene annotations, a transcriptome atlas, and signatures of genome organization. The Plant Journal 93:338–354.
Sorghum bicolor is a drought tolerant C4 grass used for the production of grain, forage, sugar, and lignocellulosic biomass and a genetic model for C4 grasses due to its relatively small genome (approximately 800 Mbp), diploid genetics, diverse germplasm, and colinearity with other C4 grass genomes. In this study, deep sequencing, genetic linkage analysis, and transcriptome data were used to produce and annotate a high‐quality reference genome sequence.
Inhibition of microbial biofuel production in drought-stressed switchgrass hydrolysate
R.G. Ong et al. (2016). Inhibition of microbial biofuel production in drought-stressed switchgrass hydrolysate. Biotechnology for Biofuels 9:237.
In order to investigate the impact of interannual climate variability on biofuel production, corn stover and switchgrass were collected during 3 years with significantly different precipitation profiles, representing a major drought year (2012) and 2 years with an average precipitation for the entire season (2010 and 2013).
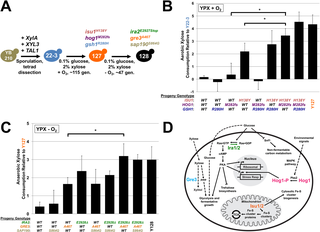
Directed Evolution Reveals Unexpected Epistatic Interactions That Alter Metabolic Regulation and Enable Anaerobic Xylose Use by Saccharomyces cerevisiae
T.K. Sato et al. (2016). Directed Evolution Reveals Unexpected Epistatic Interactions That Alter Metabolic Regulation and Enable Anaerobic Xylose Use by Saccharomyces cerevisiae. PLOS Genetics 12: e1006372
Here we combined genome sequencing, proteomic profiling, and metabolomic analyses to identify and characterize the responsible mutations in a series of evolved strains capable of metabolizing xylose aerobically or anaerobically.
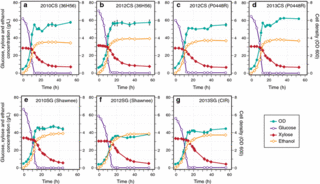
Complex Physiology and Compound Stress Responses during Fermentation of Alkali-Pretreated Corn Stover Hydrolysate by an Escherichia coli Ethanologen
M.S. Schwalbach et al. (2012). Complex Physiology and Compound Stress Responses during Fermentation of Alkali-Pretreated Corn Stover Hydrolysate by an Escherichia coli Ethanologen. Applied and Environmental Microbiology 78:3442–3457.
To gain insight into how E. coli responds to such hydrolysates, we studied an E. coli K-12 ethanologen fermenting a hydrolysate prepared from corn stover pretreated by ammonia fiber expansion.