Modeling Zymomonas Mobilis membrane interactions with lignocellulosic inhibitors
Background/Objectives
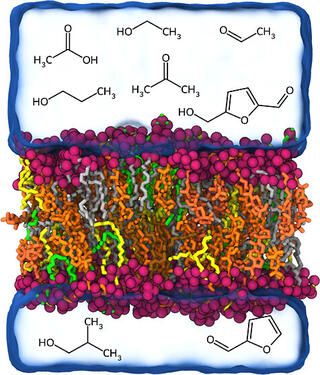
Bioenergy crops can be deconstructed using physical or chemical processes into small molecules for biological conversion to biofuels and products. Zymomonas Mobilis is one such microbial chassis that is resistant to ethanol stress but may be inhibited by the organic acids, aldehydes, alcohols, ketones, and amides present in processed biomass. One hypothesis is that these molecules interact with or disrupt the bacterial membrane, triggering stress response and hindering growth, but the exact mechanism remains unclear, hindering directed bioengineering efforts.
Approach
All-atom molecular dynamics simulations were used to investigate the interactions of Z. Mobilis membrane models with key inhibitors, including ethanol, furfural, HMF, and acetic acid. Simulations were conducted across a range of inhibitor concentrations analyzing membrane properties such as area per lipid, membrane thickness, lipid order parameter, lateral diffusion coefficient, and permeability coefficient.
Results
Generally membranes become thinner, with a higher area per lipid and lower order parameter as small molecule concentration increases. Trends were stronger for more hydrophobic molecules like isobutanol, propanol, and propanoic acid, which showed greater membrane perturbation as concentrations increased compared to other molecules. Hopanoids stabilize and order the membrane to maintain structure for smaller molecules but appear insufficient for larger hydrophobic molecules like isobutanol.
Impacts
Findings provide a mechanistic understanding of how small molecules in biomass degradation streams interact with the Z. mobilis membrane, offering valuable insights for future strain engineering efforts to optimize biofuel and bioproduct synthesis from biomass. Modifying membrane composition could improve microbial survival and therefore industrial biorefinery yields.
Citation
Singh, N. K., & Vermaas, J. V. Quantifying Membrane Structure and Dynamics during Bioproduct Production in Zymomonas mobilis by Molecular Simulation. The Journal of Physical Chemistry B. (2026). [DOI:10.1021/acs.jpcb.5c06231]