Decoding Abiotic Stress Resilience in Sorghum
Background/Objective
Sorghum is a stress resilient crop and a promising biofuel candidate. Given changing global climate patterns, developing crops that can be grown in intense environments is vital to maintaining robust food and energy sources. The interplay between sorghum genes and tissue specific responses to abiotic stresses at various growth points remains underexplored.
Approach
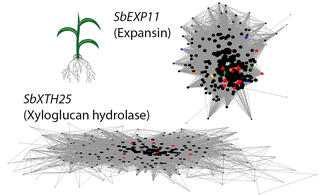
Researchers exposed sorghum plant leaves and roots to drought, heat, and salinity and performed RNA-sequenced and principal component analysis to determine if time, stress type, or tissue type was the greatest factor in gene expression. Two groups of genes similarly expressed in different conditions present (co-expression modules) in both leaves and shoots were identified, gene expression and transcription factors were compared and co-expression networks were built to analyze connectivity and gene interaction strength within each co-expression module.
Results
Tissue specific gene regulatory networks were found to be the largest force in transcriptomic changes. Genes within co-expression modules were shown to be highly connected, but transcription factors were found to be different.
Significance/Impact
This work shows co-expression networks to be a valuable tool in determining the interconnectedness of genes and in identifying genetic hubs related to stress. This strategy could be useful in identifying the tissue specific regulatory hub genes necessary to develop climate-resilient biofuel crops without sacrificing growth.
Ko, D. K., & Brandizzi, F. A tissue-resolved, network-based transcriptomic framework for abiotic stress responses in sorghum. The Plant Journal, 126, e70834. (2026). [DOI:10.1111/tpj.70834]