Dual approaches help scientists pinpoint genes that help microbes tolerate stressors
The Science
Microbes can produce biofuels and other products from plants, but production is often limited by toxic chemicals produced by plants or the process of breaking them down to separate sugars for microbial fermentation. Genetic modification of the microbes can often overcome these limitations, but without a map, it's difficult to identify which of the thousands of genes will give the cells an advantage. Traditionally scientists grow “libraries” of mutants — pools of cells with different genes turned off — in the presence of a chemical. Analyzing the surviving cells can provide clues about which genes allow the microbe to tolerate that chemical. But using only one screening method tends to produce false positives, requiring slow and labor-intensive follow-up experiments.
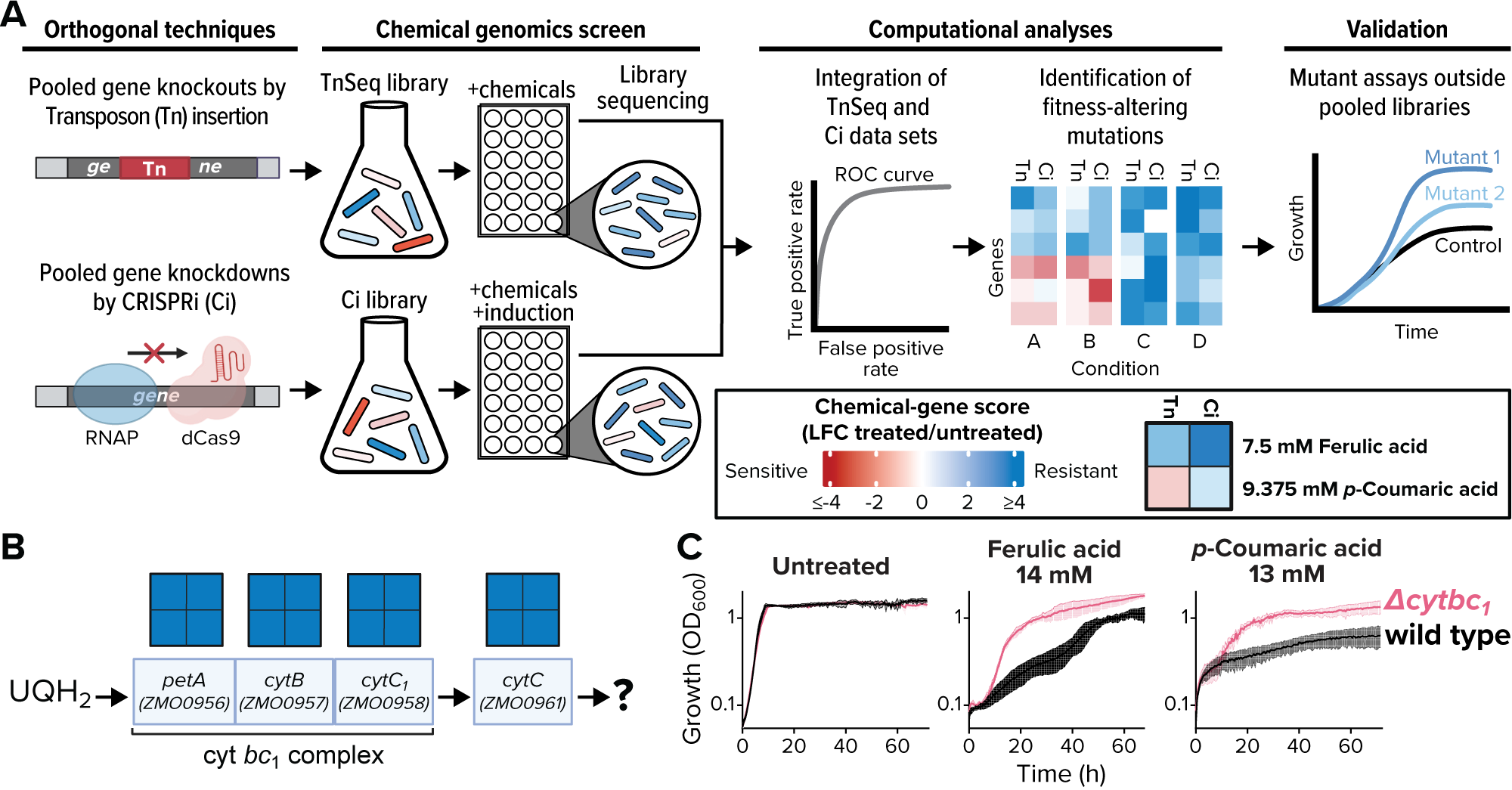
To solve the problem, scientists used two genome screening techniques with complementary strengths and weaknesses to study the effects of inhibitory chemicals on Zymomonas mobilis, a promising microbe for industrial production. Applying the dual-library approach narrowed the field of candidate genes by two thirds. Genes selected for follow-up tests were all true positives.
The Impact
Engineering microbes to tolerate chemical stressors can increase yields of biofuels and other plant-based chemicals. In addition to identifying target genes that improve Z. mobilis' resistance to toxic chemicals found in deconstructed plant material, this work provides a strategy that can be broadly applied to understanding microbial responses to chemicals across systems, paving the way for advances in biotechnology, medicine, and agriculture.
Summary
Scientists within the Great Lakes Bioenergy Research Center screened genome-scale CRISPRi (CRISPR interference) and TnSeq (transposon insertion sequencing) libraries of the Alphaproteobacterium Zymomonas mobilis against growth inhibitors commonly found in deconstructed plant biomass. By integrating data from the two techniques, they narrowed the genetic search space from 103 genes identified by at least one library to 31 cross-validated targets. A subset of those were validated using precise gene deletions, indicating the strategy precisely identifies true positives.
The experiment identified all known genes in the cytochrome bc1 and cytochrome c synthesis pathway as potential targets for engineering resistance to phenolic acids under anaerobic conditions, a subset of which were validated using precise gene deletions. This finding is specific to the cytochrome bc1 and cytochrome c pathway and does not extend to other branches of the electron transport chain. Results also showed that exposure to one phenolic acid causes substantial remodeling of the cell envelope proteome in Z. mobilis as well as the downregulation of TonB-dependent transporters.