Ancient co-option of LTR retrotransposons as yeast centromeres
M.A.B. Haase et al. "Ancient co-option of LTR retrotransposons as yeast centromeres" Nature (2026) [DOI:10.1038/s41586-025-10092-0]
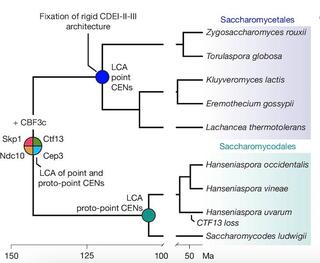
Centromeres ensure accurate chromosome segregation, yet their DNA evolves rapidly across eukaryotes leaving the origins of new centromere architectures unclear1,2,3,4. The brewer’s yeast Saccharomyces cerevisiae exemplifies this long-standing puzzle. Its centromeres shifted ancestrally from large, repeat-rich, epigenetically specified forms to the compact, genetically defined ‘point’ centromeres1,5. How this transition occurred has remained unresolved6. Here we identify evolutionarily related ‘proto-point’ centromeres that provide a resolution to the evolutionary origins of point centromeres. Proto-point centromeres contain a single centromeric nucleosome positioned over an AT-rich core, accompanied by relaxed organization and sequence variability of flanking cis-elements. In two species, these proto-point centromeres lie within retrotransposon-derived repeat clusters, linking ancestral repeat-rich centromeres to genetically encoded ones. Comparative and phylogenetic analyses indicate that proto-point and point centromeres evolved in an ancestor with retrotransposon-rich centromeres. These results identify long-terminal-repeat retrotransposons, specifically Ty5 sequences, as the genetic substrate for point-centromere evolution and provide a mechanistic route by which an epigenetic centromere can become genetically specified. More broadly, they show how selfish elements can be co-opted to perform essential chromosomal functions.
All yeast strains and plasmids are available on request to M.A.B.H. or J.D.B. Raw whole-genome, MNase-seq and RNA sequencing data were deposited in the SRA and are available under the BioProject ID PRJNA961205 with SRA accessions SRR32695949–SRR32695985, SRR36083336–SRR36083338 and SRR36267482–SRR36267484. Assembled genomes are available under the BioProject ID PRJNA961205 with accessions JBNIMK000000000, JBNIML000000000, JBNIMM000000000, JBSZQV000000000, JBSZQW000000000 and JBSZQX000000000. Processed MNase-seq and ChIP–seq data are available at the Gene Expression Omnibus accession GSE294411. Genome sequences were downloaded from NCBI and the accessions are listed in Supplementary Data 7. Source data are provided with this paper.